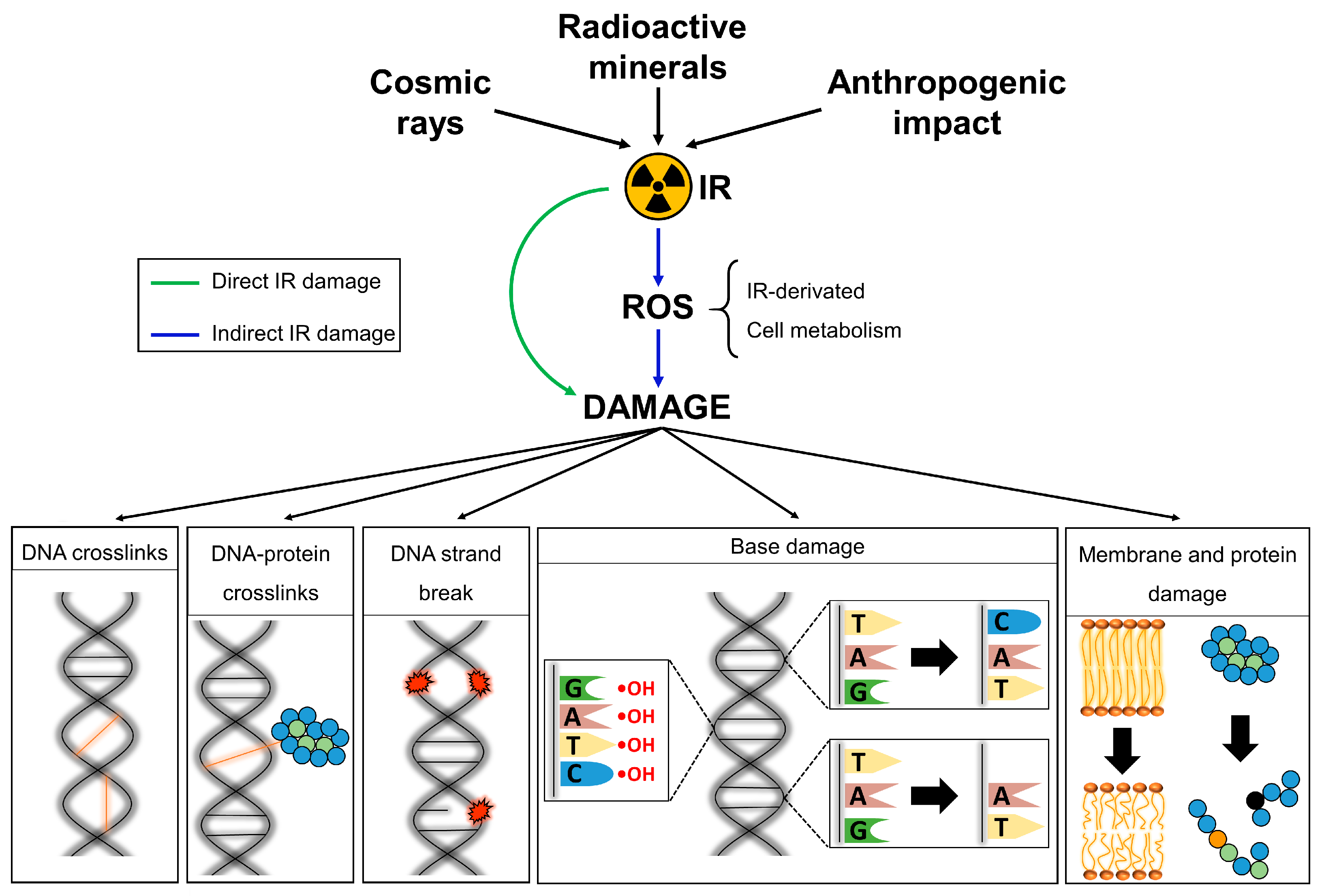
Plants | Free Full-Text | Chronic Ionizing Radiation of Plants: An Evolutionary Factor from Direct Damage to Non-Target Effects

IJMS | Free Full-Text | Cracking the Floral Quartet Code: How Do Multimers of MIKCC-Type MADS-Domain Transcription Factors Recognize Their Target Genes?

Exploring protein phosphorylation by combining computational approaches and biochemical methods - ScienceDirect

Computational methods and challenges in analyzing intratumoral microbiome data: Trends in Microbiology
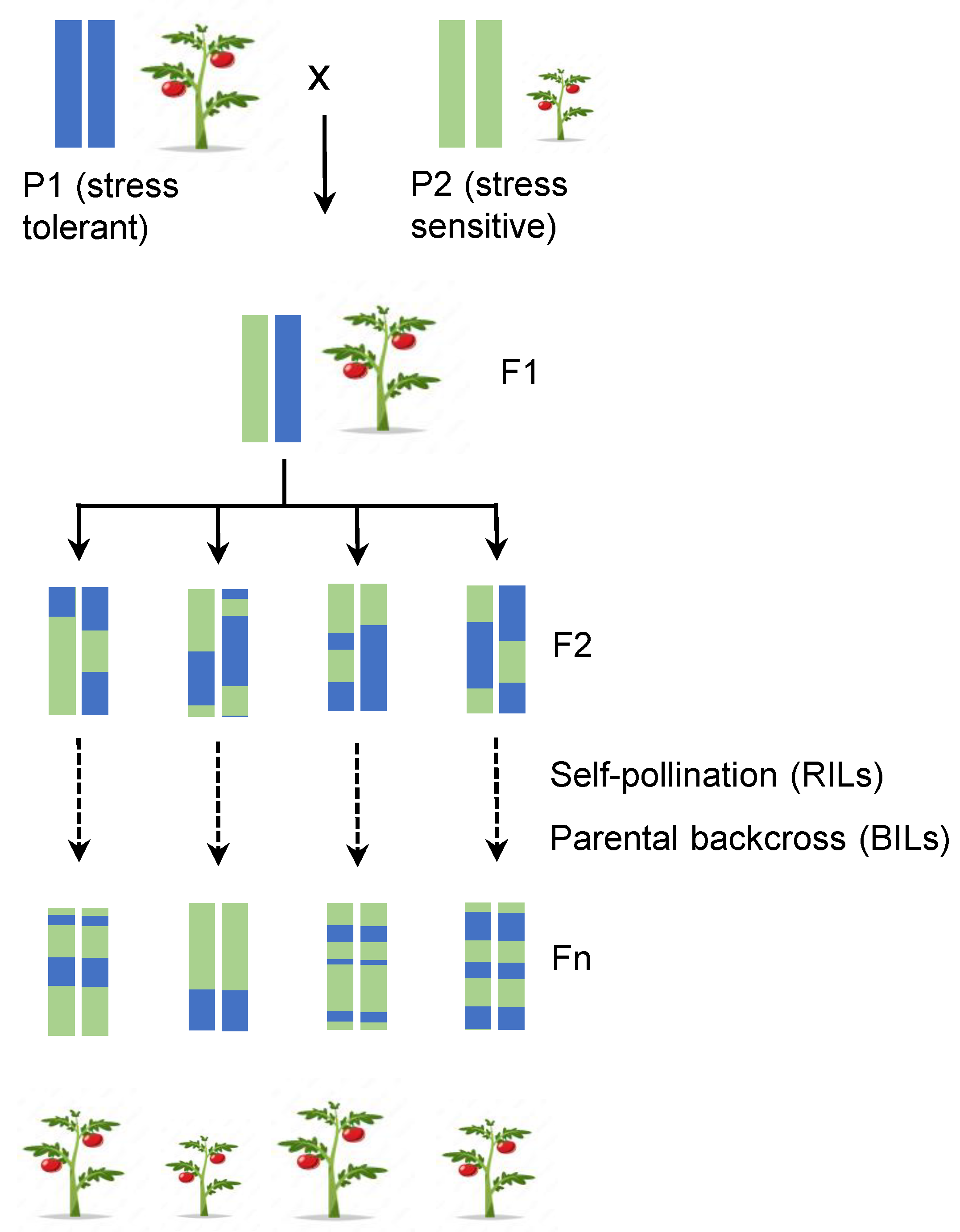
IJMS | Free Full-Text | From Classical to Modern Computational Approaches to Identify Key Genetic Regulatory Components in Plant Biology
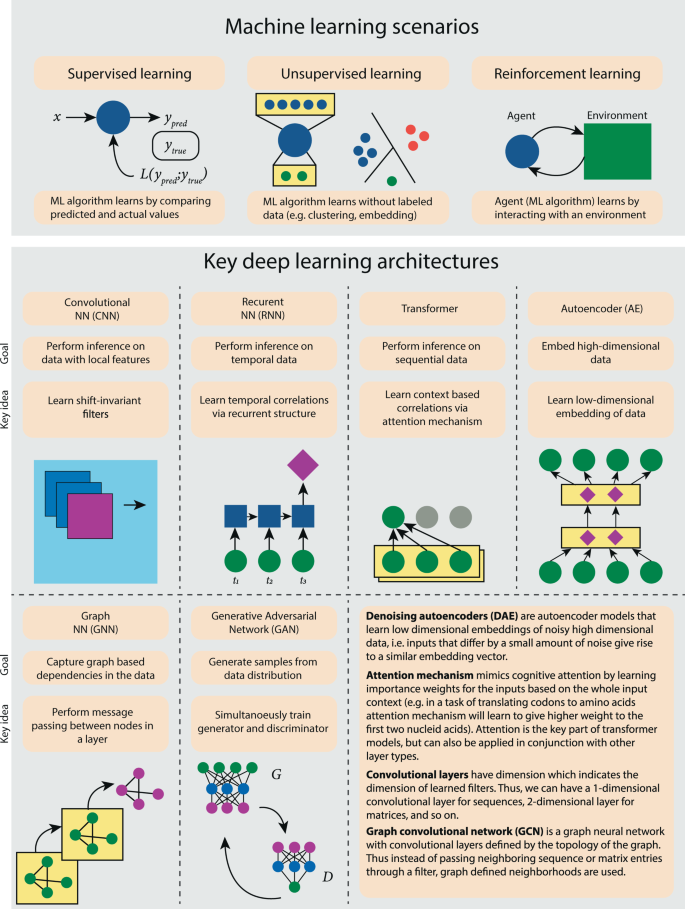
Current progress and open challenges for applying deep learning across the biosciences | Nature Communications

Frontiers | Computational Enzyme Engineering Pipelines for Optimized Production of Renewable Chemicals
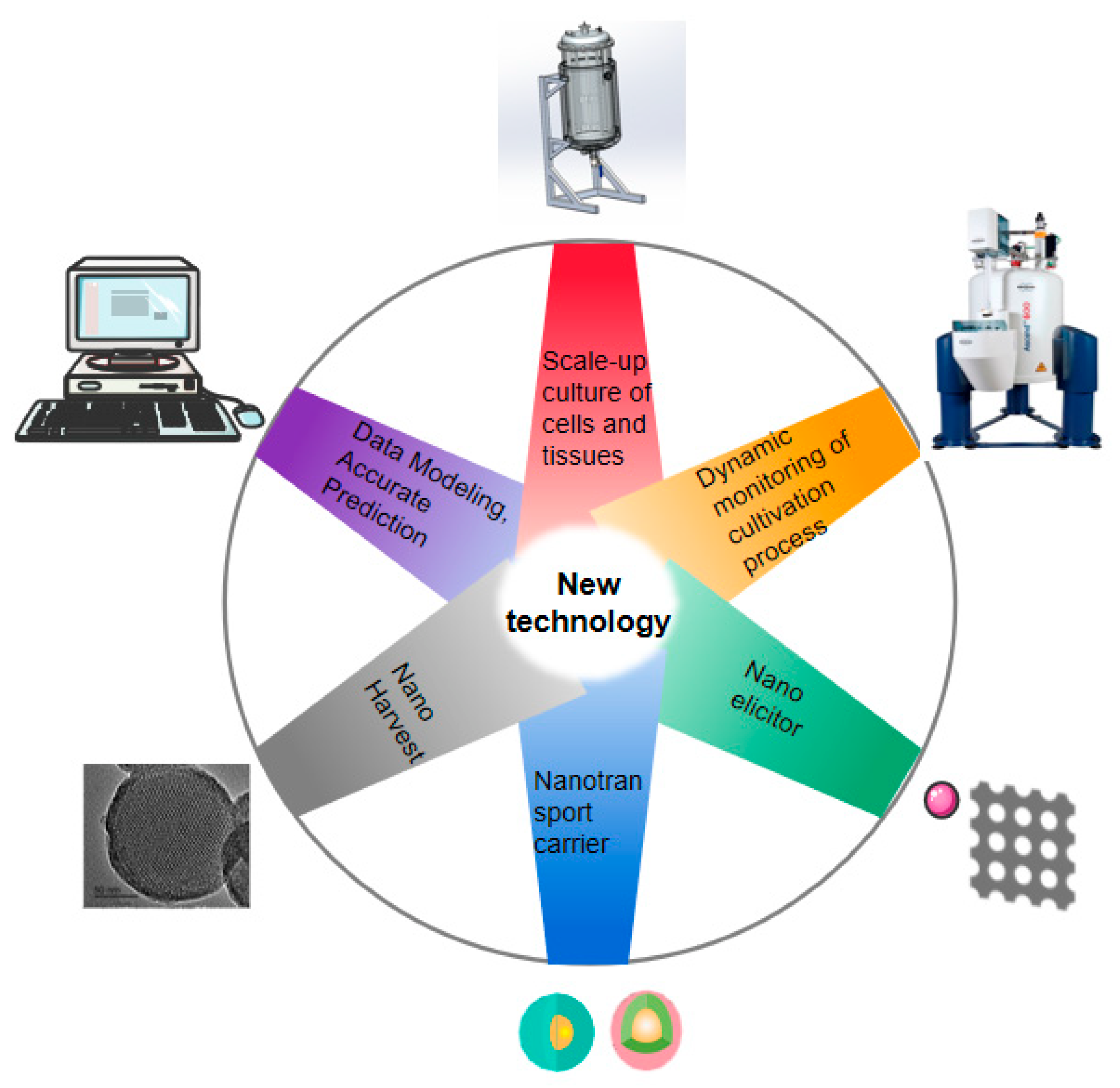
Plants | Free Full-Text | Application of Data Modeling, Instrument Engineering and Nanomaterials in Selected Medid the Scientific Recinal Plant Tissue Culture

A computational framework for identifying the transcription factors involved in enhancer-promoter loop formation: Molecular Therapy - Nucleic Acids

Progress toward rapid, at-line N-glycosylation detection and control for recombinant protein expression - ScienceDirect

Artificial intelligence for drug discovery: Resources, methods, and applications: Molecular Therapy - Nucleic Acids